Another
documentary has surfaced that leans on the apprehension or anticipation that genetics will confirm the intellectual advantages of certain racial groups over others. Realistically, I doubt Nature or The New York Times will break such a story. The media generally does not even address racial differences in the warrior gene. Why should anyone expect a mainstream science reporter to painstakingly calculate the cumulative effect of who-knows-how-many single nucleotide polymorphisms (SNPs) potentially to prove right Southern bigots? Nevertheless, curiosity abhors a pat tune, and I think questions of race naturally meld into one of the most basic existential questions: What does it mean to be human? In general, examinations of the genetics of obesity and intelligence would complement each other not only because both traits have complex genetic architectures, but also because obesity is a less controversial subject for many than intelligence, especially when these subjects intersect with race. So, an approach that gains acceptance for less contested phenotypes will streamline an IQ juggernaut. Since stepping on a scale is far simpler than measuring intelligence, temperament, personality, or behavior, that genome-wide association studies (GWAS) for body-mass index (BMI) are further along does not surprise.
So far, GWAS have
identified 32 genetic loci for obesity. Different studies have used different SNPs to represent these loci. In order to compare diverse ethnic populations at these loci, I entered each SNP into the HapMap online database. Then, I selected the SNP from each locus for which HapMap provides the most information. HapMap has very thorough data for Northern Europeans, the Yoruba of Africa, Chinese people, and Japanese people. By multiplying the respective effect sizes of each SNP by each group’s allele frequency and adding the results for each group, I could graph a genetic index of obesity for each of those four groups. I also added the data from those four groups to data from less represented ethnic groups to create the following broader racial or ethnic designations. “Black” refers to the Yoruba, the Luhya, the Maasai, and African Americans. “Whites” are Northern Europeans and Italians. “East Asians” are Chinese and Japanese people, and the group “Asians” also adds people from India. The resulting graph suggests that Asians have a lower genetic risk for obesity.

For a more detailed picture of the full range of ethnic groups, I removed 7 of the 32 loci that had more limited data. This graph still seems to show less obesity propensity for Asians. In fact, graphs like this can serve as counterpoints to the social deconstruction of race, since ethnic groups within a continental racial group do tend to cluster together in allele frequencies. This fits with recent population genetics studies. For instance, a
new study of natural selection in African populations found that “positive selection does not appear to have substantially shaped present-day allele frequency differences among the African populations in our dataset…. Our results agree with Coop
et al (2009) and Pickrell
et al (2009), who found that selective sweep signals tend to cluster by broad geographic and continental regions…”
Perhaps the similarity of genetic risk for white and black people should not surprise.
Currently, in the United States, adult black women have nearly twice the prevalence of obesity as adult white women, but for the men no statistically significant difference exists. Therefore, I suspect that the unfortunate obesity epidemic among African-American women is a cultural phenomenon, rather than genetic destiny.
A relevant criticism of my genetic racial comparisons is that the GWAS that identified these genes were conducted in Europeans. Moreover, Chinese people have allele frequencies of zero for 5 of the loci, and Japanese people have allele frequencies of zero for those 5 and one more. If those loci would not be identified in Chinese or Japanese obesity GWAS, one could certainly imagine that those GWAS could identify obesity-causing alleles which whites or Africans lack. Therefore, I recreated the first graph minus those 6 loci to attempt a more fair comparison.
The racial genetic risk gap is lessened but is still very much present.
A different set of five loci (four for the detailed ethnicity breakdown graph) affect extreme obesity risk, with extreme obesity defined as an adult BMI of greater than or equal to 40 or a childhood BMI greater than or equal to 99 percent of the age and gender cohort. In the case of extreme obesity, Asians appear to be at greater risk than whites. Japanese people, in particular, apparently possess a sumo-sized extreme obesity risk, despite having low overall genetic obesity risk.


Three SNPs affect body fat composition, as measured by bioimpedance analysis and dual energy X-ray absorptiometry. One of the alleles is a member of the 32 obesity loci. Another was found to affect body fat percentage in Europeans but not Indians. The third, IRS1, has an allele that raises body fat but paradoxically lowers type 2 diabetes risk in men, seemingly by shifting fat storage to the layer just beneath the skin where it is less harmful. Asians are much less likely to have that allele, which could help explain why
studies are finding that nonoverweight Chinese people have high rates of metabolic abnormalities more commonly associated with obesity. Specifically, one-third of nonoverweight Chinese people have at least one metabolic risk factor.
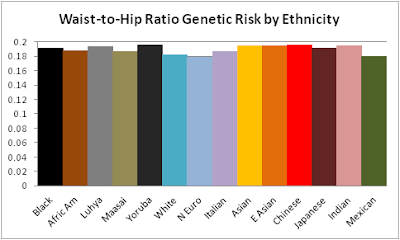
GWAS have
found fourteen SNPs so far for waist-to-hip ratio after controlling for BMI, age, and sex. The detailed ethnicity breakdown bar graph includes eleven of them. These graphs do not show strong racial or ethnic differences, but perhaps these alleles further contribute to unhealthy fat distribution in Asians.

The overriding concern that troubles this form of analysis is that the totality of the molecular genetics of any of these phenotypes is still so poorly detailed that the known loci account for almost none of the genetic heritability determined by twins studies and the like. The obesity GWAS used a quarter of a million subjects to lay out just 2 to 4 percent of the estimated heritability. The GWAS for waist-to-hip ratio used 190,000 subjects to account for 2 to 5 percent of the estimated heritability. The three body-fat SNPs using 76,000 subjects explain a mere 0.25% of body fat composition heritability. Despite such low levels of explained variance, this genetic data accurately samples the whole of which it is part.
Belsky et al recently demonstrated this to be the case, using the same method of calculating an obesity genetic risk score applied to individuals rather than groups. In fact, the effect size of the genetic risk index correlated only slightly less than familial risk based on each individual’s parents’ BMI, and their genetic risk index did not even include 3 of the 32 loci. Also, as the graphs below reveal, the genetic risk was not merely a subset of the parental risk. The two risk scores (listed as high or low for being one standard deviation above or below the mean, respectively) could not completely match the predictive quality of a risk based on the two in combination.



Presumably individual and population differences in important characteristics have some comprehensible root cause or causes. Regardless of the precise contributions to polygenic trait evolution from natural selection, the Founder effect, deleterious mutations, and so on, the order of allele identification is sufficiently independent of these forces, and the effect sizes are sufficiently distributed so as to make, I predict, nearly any genetic index a representative sample. If I am wrong, then at least I have started a scalable database as additional loci trickle in.
The Latest Intelligence on Intelligence
The concern about applicability to non-Europeans has greater salience, considering recent findings about rare SNPs. These GWAS only consider the independent effects of common SNPs, not the effects of rare SNPs or the “non-additive” genetic effects of the interactions between genes (called epistasis). A pair of studies recently addressed rare SNPs in the journal Science.
One determined that 86% of the 500,000 SNPs found with “deep sequencing” of the protein-coding exomes were “rare,” meaning that their less common allele frequency was less than 0.5%. Rare SNPs were mostly race-specific and mostly recent deleterious mutations. Among the 1,351 European Americans (EA), 65% of all of the SNPs were race-specific. Among the 1,088 African Americans (AA), the percentage was 72%. One Native American (NA) was also examined. Below is a diagram depicting the population overlap of these SNPs and a bar graph detailing the proportion that was race-specific by allele frequency.
Research into the genetics of intelligence might also face this dilemma. A
new hypothesis from Gregory Cochran suggests that deleterious mutations determine a postulated genetic component of racial IQ gaps, with the driving force being temperature’s acceleration of the mutation rate or differences in paternal age. The authors of these studies try to explain the differences with population-size dynamics. Population growth amplifies the number of the mutations or “derived alleles” present per individual. Natural selection lowers the proportion of mutations that are “functional” or “non-synonymous,” meaning that such mutations change the protein for which the DNA codes and are usually deleterious. Recent population bottlenecks, like the exodus from Africa of Eurasians, both amplify derived alleles and only allow a shorter period of natural selection for those alleles.
It turns out that deleterious mutations are more likely to be rare SNPs in African Americans than in European Americans.
Consequently, a study with a smaller sample, such as
Lohmueller et al, will tend to find a higher proportion of deleterious-to-synonymous mutations in Europeans than Africans. For just this reason, a genetic index comparison of common SNPS for intelligence along the lines of what I have done for obesity might underestimate the genetic component of the IQ gap between black people and white people, until later research with higher sample sizes take into account rare alleles.
On the other hand, the African exodus bottleneck seems to have increased homozygosity (matching pairs) of deleterious mutations in Europeans. Although Africans seem to have more deleterious mutations per person overall, perhaps their genetic diversity and the possible recessive quality of these mutations help balance out that effect. The graph below shows the number of homozygous pairs that are synonymous (S), non-synonymous (NS), possibly damaging (PO), and probably damaging (PR).
Moreover,
MacArthur et al found much higher numbers of deleterious mutations in Asians and Africans than Europeans, but detailed follow-up determined many of these to be false positives. Thus, whites and Asians each had an equal number of true loss-of-function variants per person (104). Africans still had more (122), but each group had a roughly equal number of homozygous pairs.
Another
study that postulated a significant epistatic component to heredity used an equation based on twins studies to estimate how much of the variability of different phenotypes owes to the additive effects of SNPs (and, therefore, resulting from the sort of alleles that I am tracking). The equation result was closer to zero as the influence of those effects rose. BMI between the ages of 30 and 39 was about as close to zero (-4) as “performance IQ” (5), fitness (4), and exercise participation (5), and quite closer than general IQ (-10). BMI between the ages of 20 and 29 (18) was not as close but was still the same distance from zero as verbal IQ (-18). For comparison, birth weight was -73, and having fainting spells not in response to blood was -63. Since GWAS for IQ have already found that common SNPs account for about half of its variability, which is the bulk of its heritable component, and since those equation results showed comparable results for BMI and IQ, my approach might work fairly well for both obesity and intelligence. Nevertheless, I cannot yet reconcile that with the research on exome rare SNPs.
Steve Hsu, who recently became the Michigan State University vice president for research and graduate studies, is working on an IQ GWAS and has offered some amazing revelations in a
recent presentation. He appears to endorse deleterious mutations as the major genetic contribution to individual IQ differences, and he estimates that having 39 such mutations lowers IQ 15 points (one standard deviation), about 10,000 IQ SNPs exist, and removal of such mutations could raise IQ perhaps as much as 30 standard deviations. If today’s geniuses have IQs above about 145, one can hardly imagine the potential of a person whose IQ is over 500. Of course, no IQ test today could verify such a level, but after humanity creates those dorks, maybe they could invent one. Hsu
points to embryo selection as a realistic means of consumer-driven eugenics. He seems to think that Asian societies might be amenable to this approach, but he hopes that “progressive governments will make this procedure free for everyone.” Perhaps his work with the Beijing Genomics Institute will help identify IQ SNPs specific to Asians.
Society is used to a somewhat sporadic quality to genius because extraordinary intelligence often benefits from favorable epistasis, but embryo selection would raise the “additive” IQ potential, so the children and grandchildren of these people would invariably be super-geniuses, as well. I could conjecture about the implications of this form of eugenics. The potential to spread genius far and wide could negate a key reactionary theme, while bolstering a liberal intellectual elite. “Elite” might become a tenuous term, as genius might no longer incur reward and professional status, that is, if embryo selection becomes ubiquitous. Such circumstances multiply leftist agitants. That this might occur concomitant with global warming and automation’s realization of Marx’s “labor-saving devices” prophecy could precipitate a re-birth of Communism. At least initially, willingness to abort many healthy embryos will be a major determinant of participation, making for a far-left leading edge.
Then again, when living embodiments of eugenics ideology make quaint the quasi-religious adjective, “gifted,” an entire right-wing historical narrative will march to the fore. Intelligence will become a choice of responsible parents, and liberals will grow frustrated in their attempts to invite evolution-disbelieving African Americans.
For some, other races will always be “the other,” regardless of IQ, and universal genius promotion might sooner reach out to the family-pet community. Don’t laugh! If humans can reach a new brilliance beyond superlatives, who could say how far into the animal kingdom human intellect could penetrate? If scientists can resurrect Neandertals or Denisovans, those creatures might even share some of the target IQ variants with modern humans. All this existential tumult will owe to a movement that began with elites’ innumerable abortions. Personhood will never be the same.
 Belsky DW, Moffitt TE, Houts R, Bennett GG, Biddle AK, Blumenthal JA, Evans JP, Harrington H, Sugden K, Williams B, Poulton R, & Caspi A (2012). Polygenic risk, rapid childhood growth, and the development of obesity: evidence from a 4-decade longitudinal study. Archives of pediatrics & adolescent medicine, 166 (6), 515-21 PMID: 22665028
Flegal KM, Carroll MD, Kit BK, & Ogden CL (2012). Prevalence of obesity and trends in the distribution of body mass index among US adults, 1999-2010. JAMA : the journal of the American Medical Association, 307 (5), 491-7 PMID: 22253363
Gordon-Larsen P, Adair LS, Meigs JB, Mayer-Davis E, Herring A, Yan S, Zhang B, Shufa D, & Popkin BM (2012). Discordant Risk: Overweight and cardiometabolic risk in Chinese adults Gordon-Larsen: Overweight and Cardiometabolic Risk in China. Obesity (Silver Spring, Md.) PMID: 22714089
Granka, JM, Henn, BM, Gignoux, CR, Kidd, JM, Bustamante, CD, & Feldman, MW (2012). Limited Evidence for Classic Selective Sweeps in African Populations Genetics DOI: 10.1534/genetics.112.144071
Heid IM, Jackson AU, Randall JC, Winkler TW, Qi L, Steinthorsdottir V, Thorleifsson G, Zillikens MC, Speliotes EK, Mägi R, Workalemahu T, White CC, Bouatia-Naji N, Harris TB, Berndt SI, Ingelsson E, Willer CJ, Weedon MN, Luan J, Vedantam S, Esko T, Kilpeläinen TO, Kutalik Z, Li S, Monda KL, Dixon AL, Holmes CC, Kaplan LM, Liang L, Min JL, Moffatt MF, Molony C, Nicholson G, Schadt EE, Zondervan KT, Feitosa MF, Ferreira T, Lango Allen H, Weyant RJ, Wheeler E, Wood AR, MAGIC, Estrada K, Goddard ME, Lettre G, Mangino M, Nyholt DR, Purcell S, Smith AV, Visscher PM, Yang J, McCarroll SA, Nemesh J, Voight BF, Absher D, Amin N, Aspelund T, Coin L, Glazer NL, Hayward C, Heard-Costa NL, Hottenga JJ, Johansson A, Johnson T, Kaakinen M, Kapur K, Ketkar S, Knowles JW, Kraft P, Kraja AT, Lamina C, Leitzmann MF, McKnight B, Morris AP, Ong KK, Perry JR, Peters MJ, Polasek O, Prokopenko I, Rayner NW, Ripatti S, Rivadeneira F, Robertson NR, Sanna S, Sovio U, Surakka I, Teumer A, van Wingerden S, Vitart V, Zhao JH, Cavalcanti-Proença C, Chines PS, Fisher E, Kulzer JR, Lecoeur C, Narisu N, Sandholt C, Scott LJ, Silander K, Stark K, Tammesoo ML, Teslovich TM, Timpson NJ, Watanabe RM, Welch R, Chasman DI, Cooper MN, Jansson JO, Kettunen J, Lawrence RW, Pellikka N, Perola M, Vandenput L, Alavere H, Almgren P, Atwood LD, Bennett AJ, Biffar R, Bonnycastle LL, Bornstein SR, Buchanan TA, Campbell H, Day IN, Dei M, Dörr M, Elliott P, Erdos MR, Eriksson JG, Freimer NB, Fu M, Gaget S, Geus EJ, Gjesing AP, Grallert H, Grässler J, Groves CJ, Guiducci C, Hartikainen AL, Hassanali N, Havulinna AS, Herzig KH, Hicks AA, Hui J, Igl W, Jousilahti P, Jula A, Kajantie E, Kinnunen L, Kolcic I, Koskinen S, Kovacs P, Kroemer HK, Krzelj V, Kuusisto J, Kvaloy K, Laitinen J, Lantieri O, Lathrop GM, Lokki ML, Luben RN, Ludwig B, McArdle WL, McCarthy A, Morken MA, Nelis M, Neville MJ, Paré G, Parker AN, Peden JF, Pichler I, Pietiläinen KH, Platou CG, Pouta A, Ridderstråle M, Samani NJ, Saramies J, Sinisalo J, Smit JH, Strawbridge RJ, Stringham HM, Swift AJ, Teder-Laving M, Thomson B, Usala G, van Meurs JB, van Ommen GJ, Vatin V, Volpato CB, Wallaschofski H, Walters GB, Widen E, Wild SH, Willemsen G, Witte DR, Zgaga L, Zitting P, Beilby JP, James AL, Kähönen M, Lehtimäki T, Nieminen MS, Ohlsson C, Palmer LJ, Raitakari O, Ridker PM, Stumvoll M, Tönjes A, Viikari J, Balkau B, Ben-Shlomo Y, Bergman RN, Boeing H, Smith GD, Ebrahim S, Froguel P, Hansen T, Hengstenberg C, Hveem K, Isomaa B, Jørgensen T, Karpe F, Khaw KT, Laakso M, Lawlor DA, Marre M, Meitinger T, Metspalu A, Midthjell K, Pedersen O, Salomaa V, Schwarz PE, Tuomi T, Tuomilehto J, Valle TT, Wareham NJ, Arnold AM, Beckmann JS, Bergmann S, Boerwinkle E, Boomsma DI, Caulfield MJ, Collins FS, Eiriksdottir G, Gudnason V, Gyllensten U, Hamsten A, Hattersley AT, Hofman A, Hu FB, Illig T, Iribarren C, Jarvelin MR, Kao WH, Kaprio J, Launer LJ, Munroe PB, Oostra B, Penninx BW, Pramstaller PP, Psaty BM, Quertermous T, Rissanen A, Rudan I, Shuldiner AR, Soranzo N, Spector TD, Syvanen AC, Uda M, Uitterlinden A, Völzke H, Vollenweider P, Wilson JF, Witteman JC, Wright AF, Abecasis GR, Boehnke M, Borecki IB, Deloukas P, Frayling TM, Groop LC, Haritunians T, Hunter DJ, Kaplan RC, North KE, O'Connell JR, Peltonen L, Schlessinger D, Strachan DP, Hirschhorn JN, Assimes TL, Wichmann HE, Thorsteinsdottir U, van Duijn CM, Stefansson K, Cupples LA, Loos RJ, Barroso I, McCarthy MI, Fox CS, Mohlke KL, & Lindgren CM (2010). Meta-analysis identifies 13 new loci associated with waist-hip ratio and reveals sexual dimorphism in the genetic basis of fat distribution. Nature genetics, 42 (11), 949-60 PMID: 20935629
Hsu FC, Lenchik L, Nicklas BJ, Lohman K, Register TC, Mychaleckyj J, Langefeld CD, Freedman BI, Bowden DW, & Carr JJ (2005). Heritability of body composition measured by DXA in the diabetes heart study. Obesity research, 13 (2), 312-9 PMID: 15800289
Kilpeläinen TO, Zillikens MC, Stančákova A, Finucane FM, Ried JS, Langenberg C, Zhang W, Beckmann JS, Luan J, Vandenput L, Styrkarsdottir U, Zhou Y, Smith AV, Zhao JH, Amin N, Vedantam S, Shin SY, Haritunians T, Fu M, Feitosa MF, Kumari M, Halldorsson BV, Tikkanen E, Mangino M, Hayward C, Song C, Arnold AM, Aulchenko YS, Oostra BA, Campbell H, Cupples LA, Davis KE, Döring A, Eiriksdottir G, Estrada K, Fernández-Real JM, Garcia M, Gieger C, Glazer NL, Guiducci C, Hofman A, Humphries SE, Isomaa B, Jacobs LC, Jula A, Karasik D, Karlsson MK, Khaw KT, Kim LJ, Kivimäki M, Klopp N, Kühnel B, Kuusisto J, Liu Y, Ljunggren O, Lorentzon M, Luben RN, McKnight B, Mellström D, Mitchell BD, Mooser V, Moreno JM, Männistö S, O'Connell JR, Pascoe L, Peltonen L, Peral B, Perola M, Psaty BM, Salomaa V, Savage DB, Semple RK, Skaric-Juric T, Sigurdsson G, Song KS, Spector TD, Syvänen AC, Talmud PJ, Thorleifsson G, Thorsteinsdottir U, Uitterlinden AG, van Duijn CM, Vidal-Puig A, Wild SH, Wright AF, Clegg DJ, Schadt E, Wilson JF, Rudan I, Ripatti S, Borecki IB, Shuldiner AR, Ingelsson E, Jansson JO, Kaplan RC, Gudnason V, Harris TB, Groop L, Kiel DP, Rivadeneira F, Walker M, Barroso I, Vollenweider P, Waeber G, Chambers JC, Kooner JS, Soranzo N, Hirschhorn JN, Stefansson K, Wichmann HE, Ohlsson C, O'Rahilly S, Wareham NJ, Speliotes EK, Fox CS, Laakso M, & Loos RJ (2011). Genetic variation near IRS1 associates with reduced adiposity and an impaired metabolic profile. Nature genetics, 43 (8), 753-60 PMID: 21706003
Lohmueller KE, Indap AR, Schmidt S, Boyko AR, Hernandez RD, Hubisz MJ, Sninsky JJ, White TJ, Sunyaev SR, Nielsen R, Clark AG, & Bustamante CD (2008). Proportionally more deleterious genetic variation in European than in African populations. Nature, 451 (7181), 994-7 PMID: 18288194
MacArthur DG, Balasubramanian S, Frankish A, Huang N, Morris J, Walter K, Jostins L, Habegger L, Pickrell JK, Montgomery SB, Albers CA, Zhang ZD, Conrad DF, Lunter G, Zheng H, Ayub Q, DePristo MA, Banks E, Hu M, Handsaker RE, Rosenfeld JA, Fromer M, Jin M, Mu XJ, Khurana E, Ye K, Kay M, Saunders GI, Suner MM, Hunt T, Barnes IH, Amid C, Carvalho-Silva DR, Bignell AH, Snow C, Yngvadottir B, Bumpstead S, Cooper DN, Xue Y, Romero IG, 1000 Genomes Project Consortium, Wang J, Li Y, Gibbs RA, McCarroll SA, Dermitzakis ET, Pritchard JK, Barrett JC, Harrow J, Hurles ME, Gerstein MB, & Tyler-Smith C (2012). A systematic survey of loss-of-function variants in human protein-coding genes. Science (New York, N.Y.), 335 (6070), 823-8 PMID: 22344438
Meyre D, Delplanque J, Chèvre JC, Lecoeur C, Lobbens S, Gallina S, Durand E, Vatin V, Degraeve F, Proença C, Gaget S, Körner A, Kovacs P, Kiess W, Tichet J, Marre M, Hartikainen AL, Horber F, Potoczna N, Hercberg S, Levy-Marchal C, Pattou F, Heude B, Tauber M, McCarthy MI, Blakemore AI, Montpetit A, Polychronakos C, Weill J, Coin LJ, Asher J, Elliott P, Järvelin MR, Visvikis-Siest S, Balkau B, Sladek R, Balding D, Walley A, Dina C, & Froguel P (2009). Genome-wide association study for early-onset and morbid adult obesity identifies three new risk loci in European populations. Nature genetics, 41 (2), 157-9 PMID: 19151714
Scherag A, Dina C, Hinney A, Vatin V, Scherag S, Vogel CI, Müller TD, Grallert H, Wichmann HE, Balkau B, Heude B, Jarvelin MR, Hartikainen AL, Levy-Marchal C, Weill J, Delplanque J, Körner A, Kiess W, Kovacs P, Rayner NW, Prokopenko I, McCarthy MI, Schäfer H, Jarick I, Boeing H, Fisher E, Reinehr T, Heinrich J, Rzehak P, Berdel D, Borte M, Biebermann H, Krude H, Rosskopf D, Rimmbach C, Rief W, Fromme T, Klingenspor M, Schürmann A, Schulz N, Nöthen MM, Mühleisen TW, Erbel R, Jöckel KH, Moebus S, Boes T, Illig T, Froguel P, Hebebrand J, & Meyre D (2010). Two new Loci for body-weight regulation identified in a joint analysis of genome-wide association studies for early-onset extreme obesity in French and german study groups. PLoS genetics, 6 (4) PMID: 20421936
Speliotes EK, Willer CJ, Berndt SI, Monda KL, Thorleifsson G, Jackson AU, Lango Allen H, Lindgren CM, Luan J, Mägi R, Randall JC, Vedantam S, Winkler TW, Qi L, Workalemahu T, Heid IM, Steinthorsdottir V, Stringham HM, Weedon MN, Wheeler E, Wood AR, Ferreira T, Weyant RJ, Segrè AV, Estrada K, Liang L, Nemesh J, Park JH, Gustafsson S, Kilpeläinen TO, Yang J, Bouatia-Naji N, Esko T, Feitosa MF, Kutalik Z, Mangino M, Raychaudhuri S, Scherag A, Smith AV, Welch R, Zhao JH, Aben KK, Absher DM, Amin N, Dixon AL, Fisher E, Glazer NL, Goddard ME, Heard-Costa NL, Hoesel V, Hottenga JJ, Johansson A, Johnson T, Ketkar S, Lamina C, Li S, Moffatt MF, Myers RH, Narisu N, Perry JR, Peters MJ, Preuss M, Ripatti S, Rivadeneira F, Sandholt C, Scott LJ, Timpson NJ, Tyrer JP, van Wingerden S, Watanabe RM, White CC, Wiklund F, Barlassina C, Chasman DI, Cooper MN, Jansson JO, Lawrence RW, Pellikka N, Prokopenko I, Shi J, Thiering E, Alavere H, Alibrandi MT, Almgren P, Arnold AM, Aspelund T, Atwood LD, Balkau B, Balmforth AJ, Bennett AJ, Ben-Shlomo Y, Bergman RN, Bergmann S, Biebermann H, Blakemore AI, Boes T, Bonnycastle LL, Bornstein SR, Brown MJ, Buchanan TA, Busonero F, Campbell H, Cappuccio FP, Cavalcanti-Proença C, Chen YD, Chen CM, Chines PS, Clarke R, Coin L, Connell J, Day IN, den Heijer M, Duan J, Ebrahim S, Elliott P, Elosua R, Eiriksdottir G, Erdos MR, Eriksson JG, Facheris MF, Felix SB, Fischer-Posovszky P, Folsom AR, Friedrich N, Freimer NB, Fu M, Gaget S, Gejman PV, Geus EJ, Gieger C, Gjesing AP, Goel A, Goyette P, Grallert H, Grässler J, Greenawalt DM, Groves CJ, Gudnason V, Guiducci C, Hartikainen AL, Hassanali N, Hall AS, Havulinna AS, Hayward C, Heath AC, Hengstenberg C, Hicks AA, Hinney A, Hofman A, Homuth G, Hui J, Igl W, Iribarren C, Isomaa B, Jacobs KB, Jarick I, Jewell E, John U, Jørgensen T, Jousilahti P, Jula A, Kaakinen M, Kajantie E, Kaplan LM, Kathiresan S, Kettunen J, Kinnunen L, Knowles JW, Kolcic I, König IR, Koskinen S, Kovacs P, Kuusisto J, Kraft P, Kvaløy K, Laitinen J, Lantieri O, Lanzani C, Launer LJ, Lecoeur C, Lehtimäki T, Lettre G, Liu J, Lokki ML, Lorentzon M, Luben RN, Ludwig B, MAGIC, Manunta P, Marek D, Marre M, Martin NG, McArdle WL, McCarthy A, McKnight B, Meitinger T, Melander O, Meyre D, Midthjell K, Montgomery GW, Morken MA, Morris AP, Mulic R, Ngwa JS, Nelis M, Neville MJ, Nyholt DR, O'Donnell CJ, O'Rahilly S, Ong KK, Oostra B, Paré G, Parker AN, Perola M, Pichler I, Pietiläinen KH, Platou CG, Polasek O, Pouta A, Rafelt S, Raitakari O, Rayner NW, Ridderstråle M, Rief W, Ruokonen A, Robertson NR, Rzehak P, Salomaa V, Sanders AR, Sandhu MS, Sanna S, Saramies J, Savolainen MJ, Scherag S, Schipf S, Schreiber S, Schunkert H, Silander K, Sinisalo J, Siscovick DS, Smit JH, Soranzo N, Sovio U, Stephens J, Surakka I, Swift AJ, Tammesoo ML, Tardif JC, Teder-Laving M, Teslovich TM, Thompson JR, Thomson B, Tönjes A, Tuomi T, van Meurs JB, van Ommen GJ, Vatin V, Viikari J, Visvikis-Siest S, Vitart V, Vogel CI, Voight BF, Waite LL, Wallaschofski H, Walters GB, Widen E, Wiegand S, Wild SH, Willemsen G, Witte DR, Witteman JC, Xu J, Zhang Q, Zgaga L, Ziegler A, Zitting P, Beilby JP, Farooqi IS, Hebebrand J, Huikuri HV, James AL, Kähönen M, Levinson DF, Macciardi F, Nieminen MS, Ohlsson C, Palmer LJ, Ridker PM, Stumvoll M, Beckmann JS, Boeing H, Boerwinkle E, Boomsma DI, Caulfield MJ, Chanock SJ, Collins FS, Cupples LA, Smith GD, Erdmann J, Froguel P, Grönberg H, Gyllensten U, Hall P, Hansen T, Harris TB, Hattersley AT, Hayes RB, Heinrich J, Hu FB, Hveem K, Illig T, Jarvelin MR, Kaprio J, Karpe F, Khaw KT, Kiemeney LA, Krude H, Laakso M, Lawlor DA, Metspalu A, Munroe PB, Ouwehand WH, Pedersen O, Penninx BW, Peters A, Pramstaller PP, Quertermous T, Reinehr T, Rissanen A, Rudan I, Samani NJ, Schwarz PE, Shuldiner AR, Spector TD, Tuomilehto J, Uda M, Uitterlinden A, Valle TT, Wabitsch M, Waeber G, Wareham NJ, Watkins H, Procardis Consortium, Wilson JF, Wright AF, Zillikens MC, Chatterjee N, McCarroll SA, Purcell S, Schadt EE, Visscher PM, Assimes TL, Borecki IB, Deloukas P, Fox CS, Groop LC, Haritunians T, Hunter DJ, Kaplan RC, Mohlke KL, O'Connell JR, Peltonen L, Schlessinger D, Strachan DP, van Duijn CM, Wichmann HE, Frayling TM, Thorsteinsdottir U, Abecasis GR, Barroso I, Boehnke M, Stefansson K, North KE, McCarthy MI, Hirschhorn JN, Ingelsson E, & Loos RJ (2010). Association analyses of 249,796 individuals reveal 18 new loci associated with body mass index. Nature genetics, 42 (11), 937-48 PMID: 20935630
Tennessen JA, Bigham AW, O'Connor TD, Fu W, Kenny EE, Gravel S, McGee S, Do R, Liu X, Jun G, Kang HM, Jordan D, Leal SM, Gabriel S, Rieder MJ, Abecasis G, Altshuler D, Nickerson DA, Boerwinkle E, Sunyaev S, Bustamante CD, Bamshad MJ, Akey JM, Broad GO, Seattle GO, & NHLBI Exome Sequencing Project (2012). Evolution and functional impact of rare coding variation from deep sequencing of human exomes. Science (New York, N.Y.), 337 (6090), 64-9 PMID: 22604720
Zuk O, Hechter E, Sunyaev SR, & Lander ES (2012). The mystery of missing heritability: Genetic interactions create phantom heritability. Proceedings of the National Academy of Sciences of the United States of America, 109 (4), 1193-8 PMID: 22223662
Belsky DW, Moffitt TE, Houts R, Bennett GG, Biddle AK, Blumenthal JA, Evans JP, Harrington H, Sugden K, Williams B, Poulton R, & Caspi A (2012). Polygenic risk, rapid childhood growth, and the development of obesity: evidence from a 4-decade longitudinal study. Archives of pediatrics & adolescent medicine, 166 (6), 515-21 PMID: 22665028
Flegal KM, Carroll MD, Kit BK, & Ogden CL (2012). Prevalence of obesity and trends in the distribution of body mass index among US adults, 1999-2010. JAMA : the journal of the American Medical Association, 307 (5), 491-7 PMID: 22253363
Gordon-Larsen P, Adair LS, Meigs JB, Mayer-Davis E, Herring A, Yan S, Zhang B, Shufa D, & Popkin BM (2012). Discordant Risk: Overweight and cardiometabolic risk in Chinese adults Gordon-Larsen: Overweight and Cardiometabolic Risk in China. Obesity (Silver Spring, Md.) PMID: 22714089
Granka, JM, Henn, BM, Gignoux, CR, Kidd, JM, Bustamante, CD, & Feldman, MW (2012). Limited Evidence for Classic Selective Sweeps in African Populations Genetics DOI: 10.1534/genetics.112.144071
Heid IM, Jackson AU, Randall JC, Winkler TW, Qi L, Steinthorsdottir V, Thorleifsson G, Zillikens MC, Speliotes EK, Mägi R, Workalemahu T, White CC, Bouatia-Naji N, Harris TB, Berndt SI, Ingelsson E, Willer CJ, Weedon MN, Luan J, Vedantam S, Esko T, Kilpeläinen TO, Kutalik Z, Li S, Monda KL, Dixon AL, Holmes CC, Kaplan LM, Liang L, Min JL, Moffatt MF, Molony C, Nicholson G, Schadt EE, Zondervan KT, Feitosa MF, Ferreira T, Lango Allen H, Weyant RJ, Wheeler E, Wood AR, MAGIC, Estrada K, Goddard ME, Lettre G, Mangino M, Nyholt DR, Purcell S, Smith AV, Visscher PM, Yang J, McCarroll SA, Nemesh J, Voight BF, Absher D, Amin N, Aspelund T, Coin L, Glazer NL, Hayward C, Heard-Costa NL, Hottenga JJ, Johansson A, Johnson T, Kaakinen M, Kapur K, Ketkar S, Knowles JW, Kraft P, Kraja AT, Lamina C, Leitzmann MF, McKnight B, Morris AP, Ong KK, Perry JR, Peters MJ, Polasek O, Prokopenko I, Rayner NW, Ripatti S, Rivadeneira F, Robertson NR, Sanna S, Sovio U, Surakka I, Teumer A, van Wingerden S, Vitart V, Zhao JH, Cavalcanti-Proença C, Chines PS, Fisher E, Kulzer JR, Lecoeur C, Narisu N, Sandholt C, Scott LJ, Silander K, Stark K, Tammesoo ML, Teslovich TM, Timpson NJ, Watanabe RM, Welch R, Chasman DI, Cooper MN, Jansson JO, Kettunen J, Lawrence RW, Pellikka N, Perola M, Vandenput L, Alavere H, Almgren P, Atwood LD, Bennett AJ, Biffar R, Bonnycastle LL, Bornstein SR, Buchanan TA, Campbell H, Day IN, Dei M, Dörr M, Elliott P, Erdos MR, Eriksson JG, Freimer NB, Fu M, Gaget S, Geus EJ, Gjesing AP, Grallert H, Grässler J, Groves CJ, Guiducci C, Hartikainen AL, Hassanali N, Havulinna AS, Herzig KH, Hicks AA, Hui J, Igl W, Jousilahti P, Jula A, Kajantie E, Kinnunen L, Kolcic I, Koskinen S, Kovacs P, Kroemer HK, Krzelj V, Kuusisto J, Kvaloy K, Laitinen J, Lantieri O, Lathrop GM, Lokki ML, Luben RN, Ludwig B, McArdle WL, McCarthy A, Morken MA, Nelis M, Neville MJ, Paré G, Parker AN, Peden JF, Pichler I, Pietiläinen KH, Platou CG, Pouta A, Ridderstråle M, Samani NJ, Saramies J, Sinisalo J, Smit JH, Strawbridge RJ, Stringham HM, Swift AJ, Teder-Laving M, Thomson B, Usala G, van Meurs JB, van Ommen GJ, Vatin V, Volpato CB, Wallaschofski H, Walters GB, Widen E, Wild SH, Willemsen G, Witte DR, Zgaga L, Zitting P, Beilby JP, James AL, Kähönen M, Lehtimäki T, Nieminen MS, Ohlsson C, Palmer LJ, Raitakari O, Ridker PM, Stumvoll M, Tönjes A, Viikari J, Balkau B, Ben-Shlomo Y, Bergman RN, Boeing H, Smith GD, Ebrahim S, Froguel P, Hansen T, Hengstenberg C, Hveem K, Isomaa B, Jørgensen T, Karpe F, Khaw KT, Laakso M, Lawlor DA, Marre M, Meitinger T, Metspalu A, Midthjell K, Pedersen O, Salomaa V, Schwarz PE, Tuomi T, Tuomilehto J, Valle TT, Wareham NJ, Arnold AM, Beckmann JS, Bergmann S, Boerwinkle E, Boomsma DI, Caulfield MJ, Collins FS, Eiriksdottir G, Gudnason V, Gyllensten U, Hamsten A, Hattersley AT, Hofman A, Hu FB, Illig T, Iribarren C, Jarvelin MR, Kao WH, Kaprio J, Launer LJ, Munroe PB, Oostra B, Penninx BW, Pramstaller PP, Psaty BM, Quertermous T, Rissanen A, Rudan I, Shuldiner AR, Soranzo N, Spector TD, Syvanen AC, Uda M, Uitterlinden A, Völzke H, Vollenweider P, Wilson JF, Witteman JC, Wright AF, Abecasis GR, Boehnke M, Borecki IB, Deloukas P, Frayling TM, Groop LC, Haritunians T, Hunter DJ, Kaplan RC, North KE, O'Connell JR, Peltonen L, Schlessinger D, Strachan DP, Hirschhorn JN, Assimes TL, Wichmann HE, Thorsteinsdottir U, van Duijn CM, Stefansson K, Cupples LA, Loos RJ, Barroso I, McCarthy MI, Fox CS, Mohlke KL, & Lindgren CM (2010). Meta-analysis identifies 13 new loci associated with waist-hip ratio and reveals sexual dimorphism in the genetic basis of fat distribution. Nature genetics, 42 (11), 949-60 PMID: 20935629
Hsu FC, Lenchik L, Nicklas BJ, Lohman K, Register TC, Mychaleckyj J, Langefeld CD, Freedman BI, Bowden DW, & Carr JJ (2005). Heritability of body composition measured by DXA in the diabetes heart study. Obesity research, 13 (2), 312-9 PMID: 15800289
Kilpeläinen TO, Zillikens MC, Stančákova A, Finucane FM, Ried JS, Langenberg C, Zhang W, Beckmann JS, Luan J, Vandenput L, Styrkarsdottir U, Zhou Y, Smith AV, Zhao JH, Amin N, Vedantam S, Shin SY, Haritunians T, Fu M, Feitosa MF, Kumari M, Halldorsson BV, Tikkanen E, Mangino M, Hayward C, Song C, Arnold AM, Aulchenko YS, Oostra BA, Campbell H, Cupples LA, Davis KE, Döring A, Eiriksdottir G, Estrada K, Fernández-Real JM, Garcia M, Gieger C, Glazer NL, Guiducci C, Hofman A, Humphries SE, Isomaa B, Jacobs LC, Jula A, Karasik D, Karlsson MK, Khaw KT, Kim LJ, Kivimäki M, Klopp N, Kühnel B, Kuusisto J, Liu Y, Ljunggren O, Lorentzon M, Luben RN, McKnight B, Mellström D, Mitchell BD, Mooser V, Moreno JM, Männistö S, O'Connell JR, Pascoe L, Peltonen L, Peral B, Perola M, Psaty BM, Salomaa V, Savage DB, Semple RK, Skaric-Juric T, Sigurdsson G, Song KS, Spector TD, Syvänen AC, Talmud PJ, Thorleifsson G, Thorsteinsdottir U, Uitterlinden AG, van Duijn CM, Vidal-Puig A, Wild SH, Wright AF, Clegg DJ, Schadt E, Wilson JF, Rudan I, Ripatti S, Borecki IB, Shuldiner AR, Ingelsson E, Jansson JO, Kaplan RC, Gudnason V, Harris TB, Groop L, Kiel DP, Rivadeneira F, Walker M, Barroso I, Vollenweider P, Waeber G, Chambers JC, Kooner JS, Soranzo N, Hirschhorn JN, Stefansson K, Wichmann HE, Ohlsson C, O'Rahilly S, Wareham NJ, Speliotes EK, Fox CS, Laakso M, & Loos RJ (2011). Genetic variation near IRS1 associates with reduced adiposity and an impaired metabolic profile. Nature genetics, 43 (8), 753-60 PMID: 21706003
Lohmueller KE, Indap AR, Schmidt S, Boyko AR, Hernandez RD, Hubisz MJ, Sninsky JJ, White TJ, Sunyaev SR, Nielsen R, Clark AG, & Bustamante CD (2008). Proportionally more deleterious genetic variation in European than in African populations. Nature, 451 (7181), 994-7 PMID: 18288194
MacArthur DG, Balasubramanian S, Frankish A, Huang N, Morris J, Walter K, Jostins L, Habegger L, Pickrell JK, Montgomery SB, Albers CA, Zhang ZD, Conrad DF, Lunter G, Zheng H, Ayub Q, DePristo MA, Banks E, Hu M, Handsaker RE, Rosenfeld JA, Fromer M, Jin M, Mu XJ, Khurana E, Ye K, Kay M, Saunders GI, Suner MM, Hunt T, Barnes IH, Amid C, Carvalho-Silva DR, Bignell AH, Snow C, Yngvadottir B, Bumpstead S, Cooper DN, Xue Y, Romero IG, 1000 Genomes Project Consortium, Wang J, Li Y, Gibbs RA, McCarroll SA, Dermitzakis ET, Pritchard JK, Barrett JC, Harrow J, Hurles ME, Gerstein MB, & Tyler-Smith C (2012). A systematic survey of loss-of-function variants in human protein-coding genes. Science (New York, N.Y.), 335 (6070), 823-8 PMID: 22344438
Meyre D, Delplanque J, Chèvre JC, Lecoeur C, Lobbens S, Gallina S, Durand E, Vatin V, Degraeve F, Proença C, Gaget S, Körner A, Kovacs P, Kiess W, Tichet J, Marre M, Hartikainen AL, Horber F, Potoczna N, Hercberg S, Levy-Marchal C, Pattou F, Heude B, Tauber M, McCarthy MI, Blakemore AI, Montpetit A, Polychronakos C, Weill J, Coin LJ, Asher J, Elliott P, Järvelin MR, Visvikis-Siest S, Balkau B, Sladek R, Balding D, Walley A, Dina C, & Froguel P (2009). Genome-wide association study for early-onset and morbid adult obesity identifies three new risk loci in European populations. Nature genetics, 41 (2), 157-9 PMID: 19151714
Scherag A, Dina C, Hinney A, Vatin V, Scherag S, Vogel CI, Müller TD, Grallert H, Wichmann HE, Balkau B, Heude B, Jarvelin MR, Hartikainen AL, Levy-Marchal C, Weill J, Delplanque J, Körner A, Kiess W, Kovacs P, Rayner NW, Prokopenko I, McCarthy MI, Schäfer H, Jarick I, Boeing H, Fisher E, Reinehr T, Heinrich J, Rzehak P, Berdel D, Borte M, Biebermann H, Krude H, Rosskopf D, Rimmbach C, Rief W, Fromme T, Klingenspor M, Schürmann A, Schulz N, Nöthen MM, Mühleisen TW, Erbel R, Jöckel KH, Moebus S, Boes T, Illig T, Froguel P, Hebebrand J, & Meyre D (2010). Two new Loci for body-weight regulation identified in a joint analysis of genome-wide association studies for early-onset extreme obesity in French and german study groups. PLoS genetics, 6 (4) PMID: 20421936
Speliotes EK, Willer CJ, Berndt SI, Monda KL, Thorleifsson G, Jackson AU, Lango Allen H, Lindgren CM, Luan J, Mägi R, Randall JC, Vedantam S, Winkler TW, Qi L, Workalemahu T, Heid IM, Steinthorsdottir V, Stringham HM, Weedon MN, Wheeler E, Wood AR, Ferreira T, Weyant RJ, Segrè AV, Estrada K, Liang L, Nemesh J, Park JH, Gustafsson S, Kilpeläinen TO, Yang J, Bouatia-Naji N, Esko T, Feitosa MF, Kutalik Z, Mangino M, Raychaudhuri S, Scherag A, Smith AV, Welch R, Zhao JH, Aben KK, Absher DM, Amin N, Dixon AL, Fisher E, Glazer NL, Goddard ME, Heard-Costa NL, Hoesel V, Hottenga JJ, Johansson A, Johnson T, Ketkar S, Lamina C, Li S, Moffatt MF, Myers RH, Narisu N, Perry JR, Peters MJ, Preuss M, Ripatti S, Rivadeneira F, Sandholt C, Scott LJ, Timpson NJ, Tyrer JP, van Wingerden S, Watanabe RM, White CC, Wiklund F, Barlassina C, Chasman DI, Cooper MN, Jansson JO, Lawrence RW, Pellikka N, Prokopenko I, Shi J, Thiering E, Alavere H, Alibrandi MT, Almgren P, Arnold AM, Aspelund T, Atwood LD, Balkau B, Balmforth AJ, Bennett AJ, Ben-Shlomo Y, Bergman RN, Bergmann S, Biebermann H, Blakemore AI, Boes T, Bonnycastle LL, Bornstein SR, Brown MJ, Buchanan TA, Busonero F, Campbell H, Cappuccio FP, Cavalcanti-Proença C, Chen YD, Chen CM, Chines PS, Clarke R, Coin L, Connell J, Day IN, den Heijer M, Duan J, Ebrahim S, Elliott P, Elosua R, Eiriksdottir G, Erdos MR, Eriksson JG, Facheris MF, Felix SB, Fischer-Posovszky P, Folsom AR, Friedrich N, Freimer NB, Fu M, Gaget S, Gejman PV, Geus EJ, Gieger C, Gjesing AP, Goel A, Goyette P, Grallert H, Grässler J, Greenawalt DM, Groves CJ, Gudnason V, Guiducci C, Hartikainen AL, Hassanali N, Hall AS, Havulinna AS, Hayward C, Heath AC, Hengstenberg C, Hicks AA, Hinney A, Hofman A, Homuth G, Hui J, Igl W, Iribarren C, Isomaa B, Jacobs KB, Jarick I, Jewell E, John U, Jørgensen T, Jousilahti P, Jula A, Kaakinen M, Kajantie E, Kaplan LM, Kathiresan S, Kettunen J, Kinnunen L, Knowles JW, Kolcic I, König IR, Koskinen S, Kovacs P, Kuusisto J, Kraft P, Kvaløy K, Laitinen J, Lantieri O, Lanzani C, Launer LJ, Lecoeur C, Lehtimäki T, Lettre G, Liu J, Lokki ML, Lorentzon M, Luben RN, Ludwig B, MAGIC, Manunta P, Marek D, Marre M, Martin NG, McArdle WL, McCarthy A, McKnight B, Meitinger T, Melander O, Meyre D, Midthjell K, Montgomery GW, Morken MA, Morris AP, Mulic R, Ngwa JS, Nelis M, Neville MJ, Nyholt DR, O'Donnell CJ, O'Rahilly S, Ong KK, Oostra B, Paré G, Parker AN, Perola M, Pichler I, Pietiläinen KH, Platou CG, Polasek O, Pouta A, Rafelt S, Raitakari O, Rayner NW, Ridderstråle M, Rief W, Ruokonen A, Robertson NR, Rzehak P, Salomaa V, Sanders AR, Sandhu MS, Sanna S, Saramies J, Savolainen MJ, Scherag S, Schipf S, Schreiber S, Schunkert H, Silander K, Sinisalo J, Siscovick DS, Smit JH, Soranzo N, Sovio U, Stephens J, Surakka I, Swift AJ, Tammesoo ML, Tardif JC, Teder-Laving M, Teslovich TM, Thompson JR, Thomson B, Tönjes A, Tuomi T, van Meurs JB, van Ommen GJ, Vatin V, Viikari J, Visvikis-Siest S, Vitart V, Vogel CI, Voight BF, Waite LL, Wallaschofski H, Walters GB, Widen E, Wiegand S, Wild SH, Willemsen G, Witte DR, Witteman JC, Xu J, Zhang Q, Zgaga L, Ziegler A, Zitting P, Beilby JP, Farooqi IS, Hebebrand J, Huikuri HV, James AL, Kähönen M, Levinson DF, Macciardi F, Nieminen MS, Ohlsson C, Palmer LJ, Ridker PM, Stumvoll M, Beckmann JS, Boeing H, Boerwinkle E, Boomsma DI, Caulfield MJ, Chanock SJ, Collins FS, Cupples LA, Smith GD, Erdmann J, Froguel P, Grönberg H, Gyllensten U, Hall P, Hansen T, Harris TB, Hattersley AT, Hayes RB, Heinrich J, Hu FB, Hveem K, Illig T, Jarvelin MR, Kaprio J, Karpe F, Khaw KT, Kiemeney LA, Krude H, Laakso M, Lawlor DA, Metspalu A, Munroe PB, Ouwehand WH, Pedersen O, Penninx BW, Peters A, Pramstaller PP, Quertermous T, Reinehr T, Rissanen A, Rudan I, Samani NJ, Schwarz PE, Shuldiner AR, Spector TD, Tuomilehto J, Uda M, Uitterlinden A, Valle TT, Wabitsch M, Waeber G, Wareham NJ, Watkins H, Procardis Consortium, Wilson JF, Wright AF, Zillikens MC, Chatterjee N, McCarroll SA, Purcell S, Schadt EE, Visscher PM, Assimes TL, Borecki IB, Deloukas P, Fox CS, Groop LC, Haritunians T, Hunter DJ, Kaplan RC, Mohlke KL, O'Connell JR, Peltonen L, Schlessinger D, Strachan DP, van Duijn CM, Wichmann HE, Frayling TM, Thorsteinsdottir U, Abecasis GR, Barroso I, Boehnke M, Stefansson K, North KE, McCarthy MI, Hirschhorn JN, Ingelsson E, & Loos RJ (2010). Association analyses of 249,796 individuals reveal 18 new loci associated with body mass index. Nature genetics, 42 (11), 937-48 PMID: 20935630
Tennessen JA, Bigham AW, O'Connor TD, Fu W, Kenny EE, Gravel S, McGee S, Do R, Liu X, Jun G, Kang HM, Jordan D, Leal SM, Gabriel S, Rieder MJ, Abecasis G, Altshuler D, Nickerson DA, Boerwinkle E, Sunyaev S, Bustamante CD, Bamshad MJ, Akey JM, Broad GO, Seattle GO, & NHLBI Exome Sequencing Project (2012). Evolution and functional impact of rare coding variation from deep sequencing of human exomes. Science (New York, N.Y.), 337 (6090), 64-9 PMID: 22604720
Zuk O, Hechter E, Sunyaev SR, & Lander ES (2012). The mystery of missing heritability: Genetic interactions create phantom heritability. Proceedings of the National Academy of Sciences of the United States of America, 109 (4), 1193-8 PMID: 22223662